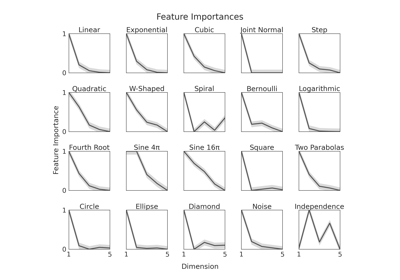
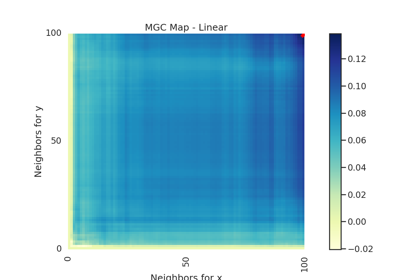
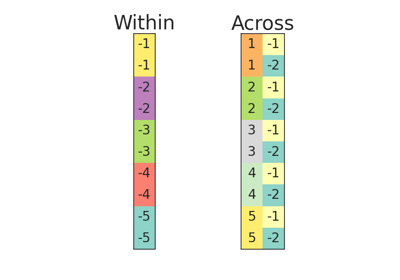
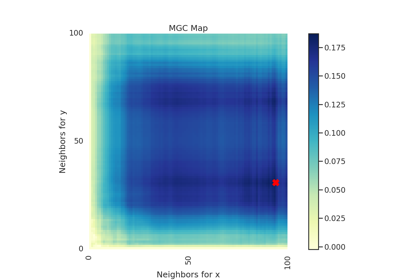
IndependenceTest¶
-
class
hyppo.independence.base.IndependenceTest(compute_distance=None, **kwargs)¶ A base class for an independence test.
- Parameters
compute_distance (
str,callable, orNone, default:"euclidean"or"gaussian") -- A function that computes the distance among the samples within each data matrix. Valid strings forcompute_distanceare, as defined insklearn.metrics.pairwise_distances,From scikit-learn: [
"euclidean","cityblock","cosine","l1","l2","manhattan"] See the documentation forscipy.spatial.distancefor details on these metrics.From scipy.spatial.distance: [
"braycurtis","canberra","chebyshev","correlation","dice","hamming","jaccard","kulsinski","mahalanobis","minkowski","rogerstanimoto","russellrao","seuclidean","sokalmichener","sokalsneath","sqeuclidean","yule"] See the documentation forscipy.spatial.distancefor details on these metrics.
Alternatively, this function computes the kernel similarity among the samples within each data matrix. Valid strings for
compute_kernelare, as defined insklearn.metrics.pairwise.pairwise_kernels,[
"additive_chi2","chi2","linear","poly","polynomial","rbf","laplacian","sigmoid","cosine"]Note
"rbf"and"gaussian"are the same metric.**kwargs -- Arbitrary keyword arguments for
compute_distkern.
Methods Summary
Calculates the independence test statistic. |
|
|
Calculates the independence test statistic and p-value. |
-
abstract
IndependenceTest.statistic(x, y)¶ Calculates the independence test statistic.
- Parameters
x,y (
ndarrayoffloat) -- Input data matrices.xandymust have the same number of samples. That is, the shapes must be(n, p)and(n, q)where n is the number of samples and p and q are the number of dimensions. Alternatively,xandycan be distance matrices, where the shapes must both be(n, n).
-
abstract
IndependenceTest.test(x, y, reps=1000, workers=1, is_distsim=True, perm_blocks=None, random_state=None)¶ Calculates the independence test statistic and p-value.
- Parameters
x,y (
ndarrayoffloat) -- Input data matrices.xandymust have the same number of samples. That is, the shapes must be(n, p)and(n, q)where n is the number of samples and p and q are the number of dimensions. Alternatively,xandycan be distance matrices, where the shapes must both be(n, n).reps (
int, default:1000) -- The number of replications used to estimate the null distribution when using the permutation test used to calculate the p-value.workers (
int, default:1) -- The number of cores to parallelize the p-value computation over. Supply-1to use all cores available to the Process.auto (
bool, default:True) -- Automatically uses fast approximation when n and size of array is greater than 20. IfTrue, and sample size is greater than 20, thenhyppo.tools.chi2_approxwill be run. Parametersrepsandworkersare irrelevant in this case. Otherwise,hyppo.tools.perm_testwill be run.is_distsim (
bool, default:True) -- Whether or notxandyare input matrices.perm_blocks (
Noneorndarray, default:None) -- Defines blocks of exchangeable samples during the permutation test. If None, all samples can be permuted with one another. Requires n rows. At each column, samples with matching column value are recursively partitioned into blocks of samples. Within each final block, samples are exchangeable. Blocks of samples from the same partition are also exchangeable between one another. If a column value is negative, that block is fixed and cannot be exchanged.
- Returns